Meet the ecologist who’s vacuuming DNA ‘out of the sky’
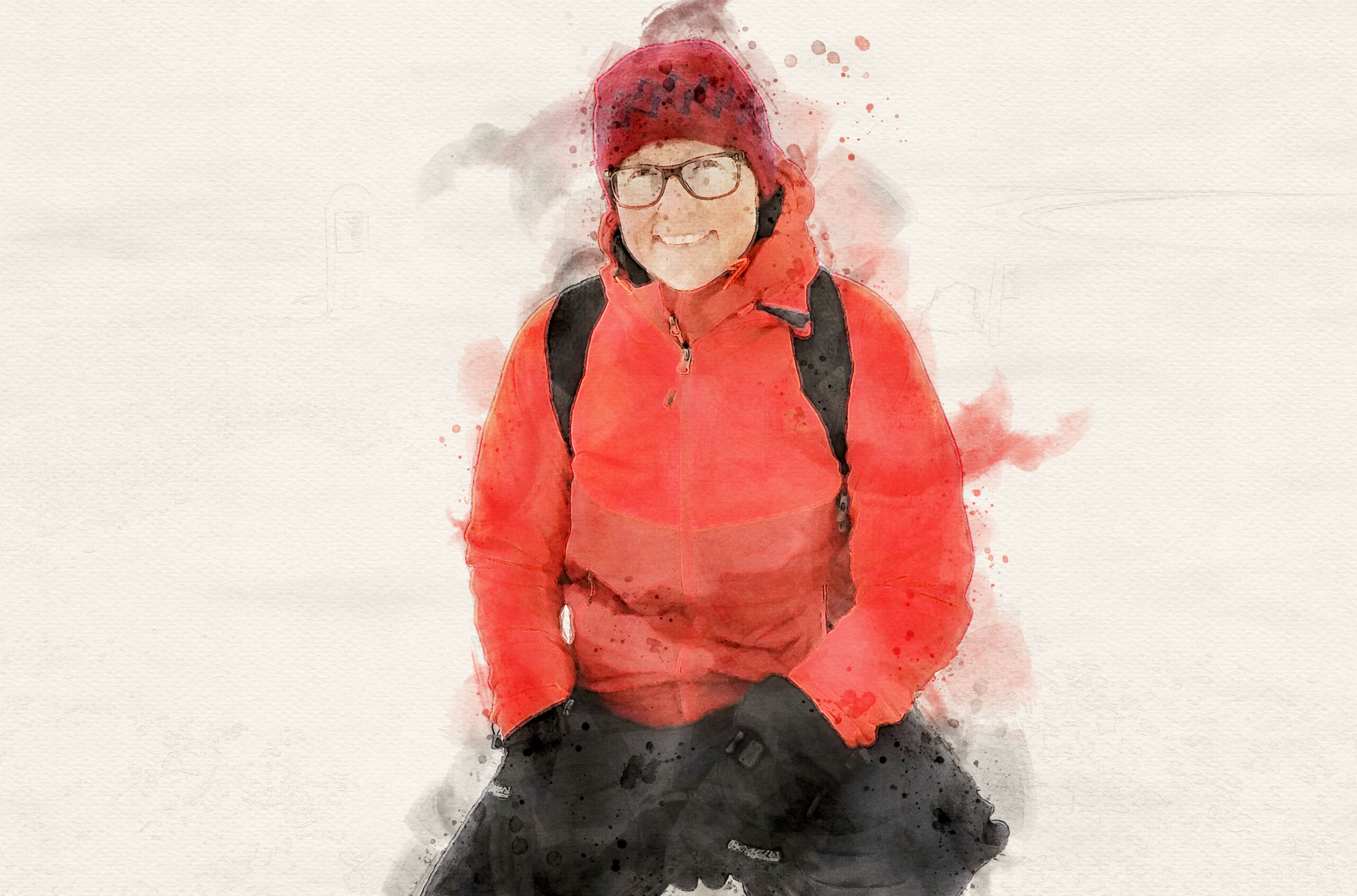
Elizabeth Clare’s work could help transform the way scientists study and monitor animal biodiversity around the world.

In December 2020, during a period of strict COVID-19 protocols, York University molecular ecologist Elizabeth Clare visited the Hamerton Zoo Park in Cambridgeshire, England. It wasn’t a pleasure outing. Instead, she was using the closed zoo as the setting for a rather outlandish experiment. “It was kind of a crazy idea. I wanted to see if we could vacuum animal DNA out of the sky,” she says.
Dr. Clare, who was working as a guest lecturer at Queen Mary University of London, set up her equipment and began collecting samples of the air from around the various animal enclosures with a small vacuum pump connected to some very fine filters. In all, her team collected 72 samples. These were then taken back to the lab and filtered onto a control paper.
A technique called polymerase chain reaction was used to amplify the amount of material collected until there was enough evidence to identify genetic markers in individual species. “It’s a bit like making coffee, but instead of pulling liquid and catching the grounds on a filter, we are vacuuming air through a filter to catch the particulate matter,” she explains.
Dr. Clare’s team identified DNA from 25 different species of animals and birds, 17 of which were known to reside at the zoo, including dingoes, gibbons, lemurs and a tiger. In the dingo enclosure, the team detected DNA from meerkats that lived 245 metres away. They even captured DNA from birds housed inside a sealed building. Unexpectedly, they also found DNA from species that did not live at the zoo, including squirrels and European hedgehogs, an endangered species in the United Kingdom.
Tracking species with environmental DNA (eDNA) is not new. Scientists have been tracking aquatic animals with DNA traces for a decade and it has also been shown that eDNA from plants can be obtained from the air. But no one had attempted to pluck animal DNA from the air until Dr. Clare, a self-professed puzzle-solver, opted to pursue this line of investigation in response to a competition launched by Queen Mary University to fund high-risk, high-impact research. “We knew DNA is found in soil and honey. We knew we could get animal DNA from the gut of a leech, so we had a good idea that there was DNA out there,” she says.
“It allows us to measure a longer-term signal. It could be very good as an early warning system for detecting the presence of invasive species or for highly sensitive species that you can’t get near, or something that is very rare and endangered.”
Dr. Clare chose a zoo to test her hypothesis because it was filled with non-native species. “If we detect tiger DNA, we know there is only one source in that environment,” she notes. The zoo was also an ideal setting because animals were housed in set locations, enabling researchers to gauge how far the DNA travelled. Even so, she admits, “We were concerned we would find nothing. I think we were all shocked at how well it worked.”
But Dr. Clare was in for another even greater shock. After assembling their data, her team put their paper online on an open-access preprint server. She was quickly inundated with emails and phone messages from people asking if she had “seen the other paper.” It turns out that Danish scientists had detected animal DNA at the Copenhagen Zoo. They employed different methods but obtained very similar results, recording the DNA of 49 animal species including rhinos, giraffes and elephants, and even picking up traces from a guppy living in a pond in the rainforest house.
It was a stunning coincidence, and the news induced some panic, because if the Danish team published first, it would seriously diminish the value of Dr. Clare’s work. She asked colleagues for advice. “There’s nothing you can do but call them,” they told her. Fortunately, Dr. Clare knew Kristine Bohmann, lead on the Danish project. They ended up striking a deal. “We decided to work together, not collaborate, but coordinate,” she says. They wrote to the journal Current Biology and sold the editors on the novel idea of publishing the two papers simultaneously in the same issue. They were published together on Jan. 6, 2022, in what Dr. Clare calls a scientific first.
“Had we not put our paper on a preprint server there probably would have been a race, and someone would have won and someone would have lost,” she says. “This way, we both won.”
Despite their success, both teams believe that the use of airborne eDNA sampling in natural environments will require more research to unlock its full potential. But they contend it could eventually transform the way scientists study and monitor animal biodiversity.
“It allows us to measure a longer-term signal. It could be very good as an early warning system for detecting the presence of invasive species or for highly sensitive species that you can’t get near, or something that is very rare and endangered,” says Dr. Clare. “It doesn’t replace other methods, but it broadens our window of detection.”
However, many questions remain unanswered. For one, the researchers still aren’t sure about the source of the eDNA they found. It could be bits of dead skin cells or fragments of hair, or saliva, or traces of urine or feces that have become aerosolized. They also don’t know why the DNA of some species escaped detection. Dr. Claire’s team missed the maned wolves, even though their urine was redolent all over the zoo, and Dr. Bohmann’s team missed the DNA of the hippos at the Copenhagen Zoo.
“We have a whole range of experiments set for this summer to answer some of these unanswered questions,” says Dr. Clare, who admits that the results of the experiment have given her a fresh outlook on her environment. “Now, when I go outside, when I go for a walk with my dog or with my kids and I take a deep breath, I think, ‘What have I just breathed in?’ It’s really traces of everything and everyone. It’s a marvelous idea that there is this information literally hanging in front of me, and if I just figure out how to catch it, then I can use it. To me that’s very exciting.”












Post a comment
University Affairs moderates all comments according to the following guidelines. If approved, comments generally appear within one business day. We may republish particularly insightful remarks in our print edition or elsewhere.
1 Comments
Great article. For more information about this field:
International Barcoding of Life – University of Guelph: the work of Dr. Paul Hebert – https://www.uoguelph.ca/ib/hebert